Jakobida
Alastair Simpson

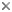
This tree diagram shows the relationships between several groups of organisms.
The root of the current tree connects the organisms featured in this tree to their containing group and the rest of the Tree of Life. The basal branching point in the tree represents the ancestor of the other groups in the tree. This ancestor diversified over time into several descendent subgroups, which are represented as internal nodes and terminal taxa to the right.

You can click on the root to travel down the Tree of Life all the way to the root of all Life, and you can click on the names of descendent subgroups to travel up the Tree of Life all the way to individual species.
For more information on ToL tree formatting, please see Interpreting the Tree or Classification. To learn more about phylogenetic trees, please visit our Phylogenetic Biology pages.
close boxIntroduction
Jakobids (Jakobida or Jakobea) are a small group of free-living, heterotrophic flagellates. There are only a dozen described species, including incertae sedis organisms, plus two undescribed species for which there are, nonetheless, substantial molecular data. Jakobids were not recognized as a group until the early 1990s, and were poorly studied until that time. Since then they have garnered considerable interest for evolutionary reasons, primarily because of their uniquely bacterial-like mitochondrial genomes.
Characteristics
Gross Features, Diversity and Habitat
Jakobids are generally less than 15 micrometres in length (mostly <10 micrometres). They have two flagella, which insert near one end of the cell (the anterior end in swimming cells). One of the two flagella can be difficult to see by light microscopy in some loricate species (see below; Petersen & Hansen, 1961, Nicholls, 1984). One flagellum is directed posteriorly and bears a single vane. The vane is very difficult to observe by light microscopy, but can be seen readily by electron microscopy (see Ultrastructure and Apomorphies). The vaned flagellum is associated with a conspicuous groove that occupies most of one side of the cell - by convention, this is regarded as the ventral side. This groove is a feeding structure – the beating action of the posterior flagellum generates a feeding current that moves suspended particles into the groove. Prokaryotes are trapped at the posterior end of the groove and are there phagocytosed.

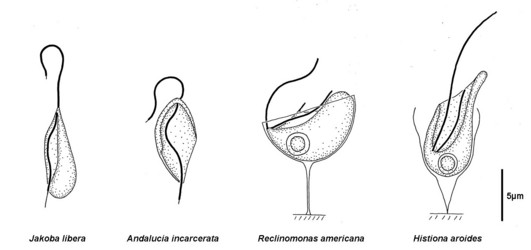
Figure 1: Some jakobids, as they appear by light microscopy. Note that the cells are drawn in different orientations - Jakoba libera is shown with the ventral side (defined by the groove) on the left. Andalucia incarcerata is shown in ventral view (looking into the ventral groove). In Reclinomonas americana the groove opens upwards (the cell 'reclines' on its dorsal side). Histiona aroides is shown roughly in ventral view. Jakoba libera is drawn in feeding position, in which the hooked portion of the anterior flagellum would adhere to the substrate. Note the loricae of Reclinomonas and Histiona, by which they attach more permanently to the substrate. © Alastair Simpson. Drawings based on light micrograph images from Patterson (1990), Simpson and Patterson (2001) and Micro*scope (by D.J. Patterson et coll.), and drawings from Nicholls (1984).
There are four formally described genera definitively assigned to Jakobids, Jakoba, Andalucia, Reclinomonas and Histiona. The latter two are grouped as the taxon Histionidae (Flavin and Nerad, 1993). Jakoba and Andalucia are free-swimming, although Jakoba libera and Andalucia incarcerata display a characteristic behaviour whereby they attach temporarily to surfaces by the distal part of the anterior flagellum (see Figure 1). This probably improves feeding efficiency. Reclinomonas and Histiona are sessile, and reside within an extracellular organic lorica. The lorica is usually shaped like a stemmed glass, with the stem (pedicel) attaching the structure to the substrate (Figure 1). During cell division the lorica is inherited by one daughter cell, while the other swims away as a zoospore, presumably subsequently settling and constructing a new lorica (O’Kelly, 1997). Cysts have been observed in several species, both loricate and aloricate (Petersen & Hansen, 1961; Mylnikov, 1984; O’Kelly, 1993, 1997; Lara et al., 2006).
Free-swimming jakobids have been recorded in a wide variety of environments, including marine, freshwater, soil and even very hypersaline habitats (Ruinen, 1938—see Patterson, 1990). One free-swimming species, Andalucia incarcerata (formerly known as Jakoba incarcerata), is a microaerophile or anaerobe (Bernard et al., 2000; Simpson and Patterson, 2001), and its mitochondrial organelle lacks cristae (see Ultrastructure and Apomorphies). By contrast, the sessile/loricate jakobids (Histionidae), including the incerte sedis organisms Stenocodon and Stomatochone (see Discussion of Phylogenetic Relationships), to our knowledge have only been recorded from freshwater habitats.
Ultrastructure and Apomorphies
The fine structure of jakobid cells is reported by Mylnikov (1989), Patterson (1990), Flavin and Nerad (1993), O’Kelly (1993, 1997), Simpson and Patterson (2001) and Lara et al. (2006). The main cellular components are mostly unremarkable: There is a single nucleus located near the flagellar bases and a single dictyosomal Golgi body, while the mitochondrial organelles have tubular cristae, except in Jakoba libera where they are flatttened, and Andalucia incarcerata, where there are no cristae (see O’Kelly, 1993; Simpson and Patterson, 2001; Lara et al., 2006). There is probably only one mitochondrion per cell in most cases (e.g. Flavin & Nerad, 1993). Histiona aroides and Reclinomonas americana have small, round extrusomes (O’Kelly, 1993, 1997), while a paranuclear microbody with single bounding membrane was reported in Andalucia godoyi (Lara et al, 2006). Electron microscopy studies show that the lorica of Reclinomonas americana has a coating of fine nail-shaped scales. These are produced intracellularly within the endomembrane system (Flavin and Nerad, 1993; O’Kelly, 1997).
The cytosketelon of jakobids has been studied in some detail, especially by Patterson (1990), O’Kelly (1997), Simpson and Patterson (2001) and Lara et al. (2006). There are two main microtubular roots in the flagellar apparatus cytoskeleton, both originating from the basal body (kinetosome) of the posterior flagellum, and both associated with a series of non-microtubular fibres (Figure 2). These flagellar roots, and microtubules derived from them, support the margins and floor of the feeding groove (Figure 3, 4). The vane on the posterior flagellum is supported by a delicate sheet of material that lies within the flagellar membrane (Figures 3 & 5). The vane is located on the dorsal side of the flagellum (i.e. the side closest to the groove floor - Figure 3).

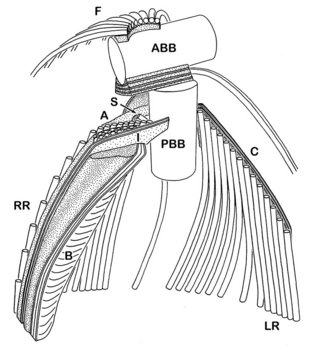
Figure 2. Andalucia incarcerata flagellar apparatus, viewed from the ventral side (i.e. the same orientation as A. incarcerata is shown in figure 1.) ABB - anterior basal body, PBB posterior basal body, LR - left microtubular root, RR - right microtubular root, S - singlet microtubule, F - Dorsal fan of microtubules, A,B,I - various non-microtubular fibres associated with the right root, C - Non microtubular fibre associated with the dorsal side of the left root. Note that only the region immediately around the flagellar bases is shown (at the top of the groove)—all of the microtubular structures extend beyond the diagram. © 2008 Alastair Simpson. Modified from Simpson and Patterson (2001).
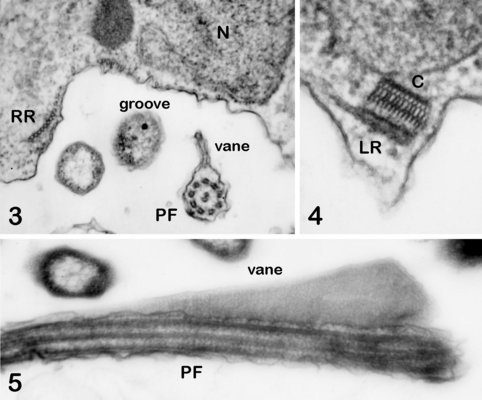
Andalucia incarcerata, transmission electron micrographs. In all figures 'ventral' is towards the bottom of the page, and dorsal towards the top. Figure 3. Transverse section of the anterior portion of the groove, showing the right microutubular root (RR) supporting the groove margin. Note the other microtubules supporting the floor of the groove. Note also the transverse section of the posterior flagellum (PF) with the single vane on its dorsal side. N: Nucleus. Figure 4. Left margin of the anterior end of the feeding groove showing a transverse section of the mult-layered 'C' fibre that supports the dorsal side of the left microtubular root (LR). The microtubules of the LR are cut through obliquely in this section, and cannot be individually distinguished. Figure 5. Longitudinal section of part of the posterior flagellum (PF) showing the proximal portion of the dorsal vane. © 2008 Alastair Simpson.
This general organization of groove and vaned flagellum is not unique to jakobids - it is seen in several other groups of Excavata, such as retortamonads, Trimastix, Malawimonas, Carpediemonas, and Dysnectes (see O’Kelly, 1993; O’Kelly and Nerad, 1999; Simpson & Patterson, 1999; Simpson, 2003; Yubuki et al., 2007). Jakobids, however, share several unique structural features relative to other excavates (see Simpson and Patterson, 2001, Simpson, 2003, Lara et al., 2006).
- The organization of the flagellar vane – single and located on the dorsal side of the flagellum (Figure 3) – is unique.
- The non-microtubular ‘C fibre’ attached to the microtubular root that supports the left margin of the groove has a distinctive multilayered substructure (Figure 4), which is different from the substructure of the homologous fibres in other Excavata (where present).
- The ‘dorsal fan’ of individual microtubules that supports the dorsal side of the cell originates almost immediately adjacent to the anterior basal body (Figure 2), rather than from along an anterior microtubular root, or in association with a distinct non-microtubular sheet that is separated from the basal body by some distance.
Mitochondrial Genomes
The mitochondrion of eukaryotes is an organelle of endosymbiotic origin, derived ultimately from a eubacterium, specifically, a member of the Alphaproteobacteria. The mitochondrial genome is the highly reduced remnant of the genome of the symbiotic bacterium. Most mitochondrial genomes are now extremely small—most animal mitochondria, for example, house just 13 protein-coding genes, plus some RNA-coding genes, and retain only faint similarities to the genomes of living prokaryotes. Amongst other things, mitochondrial genomes are not transcribed by a eubacterial-type multi-subunit (α2ββ’) RNA polymerase. The standard mitochondrial RNA polymerase is instead a single subunit enzyme that is nucleus-encoded, and is related to the RNA polymerases of T3/T7-type bacteriophage (i.e. DNA viruses).

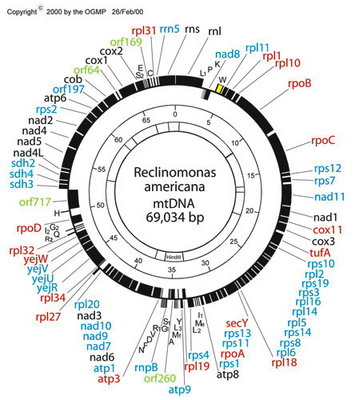
Figure 6. Map of the mitochondrial genome of the jakobid Reclinomonas americana. Red type denotes idenitified protein-coding genes that were only known from jakobid mitochondrial genomes (note that three of these are, in fact, present in Malawimonas, which is not specifically related to jakobids). The genes rpoA, rpoB, rpoC and rpoD encode subunits of eubacterial-type (α2ββ’σ) RNA polymerase. © 2000 OGMP
Jakobids constitute a glaring exception. Their mitochondrial genomes contain many more functional genes than do those of other groups of eukaryotes. The jakobid Reclinomonas americana has a mitochondrial genome that encodes nearly 100 genes in total, including some 67 protein-coding genes (Lang et al., 1997; see Figure 6). In all, ~15 of these have never been found in the mitochondrial genomes of any other group of eukaryotes (Gray et al., 2004). The ordering of genes on the Reclinomonas genome conspicuously preserves vestiges of typical proteobacterial operons (Lang et al., 1997, 1999a,b). Most protein-coding genes in Reclinomonas americana have conserved Shine-Dalgarno-like sequences upstream of the coding regions (although similar sequences do not appear to be present in Jakoba libera—Lang et al., 1999b). Most remarkably, the mitochondrial genome of Reclinomonas includes four genes that encode subunits of a classic eubacterial-type α2ββ’σ RNA polymerase (Lang et al. 1997). Other jakobid mitochondrial genomes have very similar gene complements to Reclinomonas, and also encode these four RNA polymerase subunits, except in Jakoba libera, whose mtDNA encodes only two of them (Lang et al., 1999a; Gray et al., 2004). The most straightforward interpretation is that jakobid mitochondria have retained the ancestral eubacterial RNA polymerase, which was replaced by the viral-type polymerase in all other mitochondriate eukaryotes. In summary, the mitochondrial genomes of jakobids are ‘primitively complex’—they more closely resemble the genomes of their autonomous proteobacterial ancestors than do any other mitochondria.
Discussion of Phylogenetic Relationships
Phylogenies of excavates based on morphological data recover the monophyly of jakobids with strong support (Lara et al., 2006). By contrast, the monophyly of all jakobids has been difficult to confirm in molecular phylogenetic analyses. Andalucia almost always branches separately from other jakobids in phylogenies of single genes (e.g. Edgcomb et al., 2001; Cavalier-Smith, 2003; Simpson et al., 2002, 2008; Lara et al. 2006), and of a four gene dataset (Yoon et al., 2008), while a set of analyses based on 6-7 protein-coding genes could not robustly resolve the relationships amongst Andalucia, other jakobids, Heterolobosea and Euglenozoa (Simpson et al., 2008—note that Heterolobosea and Euglenozoa are the closest relatives of jakobids—Simpson & Roger, 2004; Simpson et al., 2006; Rodriguez-Ezpeleta et al., 2007). The monophyly of jakobids is, however, strongly supported in recent phylogenomic analyses of over 100 proteins (Hampl et al., unpublished).
Within jakobids, Andalucia is monophyletic, and is the sistergroup to all other studied jakobids (Lara et al., 2006; Simpson et al., 2008, Hampl et al., unpublished). The Histionidae (~ the loricate jakobids) are generally assumed to be monophyletic, and indeed Histiona aroides and Reclinomonas americana form a clade in morphological analyses of excavates (Simpson, 2003; Lara et al., 2006) and unite as a very strongly supported group in recent phylogenomic analyses of eukaryotes (Rodriguez-Ezpeleta et al., 2007; Hampl et al., unpublished). There are, however, neither molecular nor ultrastructural data from either the other described species of Histiona, or from other taxa suspected to be histionid jakobids (Patterson et al., 2002), namely Stenocodon (which is loricate) and Stomatochone (which is sessile, but lacks a lorica—Pascher, 1943). The remaining relationships amongst jakobids are unclear—there are considerable molecular data available for Jakoha libera, and two other undescribed jakobids (referred to in the literature as ‘Seculamonas ecuadoriensis’ and ‘Jakoba bahamensis’ or ‘Jakoba bahamiensis’), but the relationships amongst these organisms and with Histionidae are not robustly resolved in published phylogenies based on several, or even >100 protein-coding genes (Rodriguez-Ezpeleta et al., 2007; Simpson et al., 2008). Notably, ‘Jakoba bahamensis’ does not form a clade with Jakoba libera in these analyses.
References
Bernard, C., Simpson, A. G. B. & Patterson, D. J. (2000) Some free-living flagellates (Protista) from anoxic habitats. Ophelia 52: 113-142.
Cavalier-Smith, T. (2003) The excavate protozoan phyla Metamonada Grasse emend. (Anaeromonadea, Parabasalia, Carpediemonas, Eopharyngia) and Loukozoa emend. (Jakobea, Malawimonas): their evolutionary affinities and new higher taxa. Int. J. Syst. Evol. Microbiol. 53: 1741-1758.
Edgcomb, V. P., Roger, A. J., Simpson, A. G. B., Kysela, D. & Sogin, M. L. (2001) Evolutionary relationships among jakobid flagellates as indicated by alpha- and beta-tubulin phylogenies. Mol. Biol. Evol. 18: 514-522.
Flavin, M. & Nerad, T. A. (1993) Reclinomonas americana n. g., n. sp., a new freshwater heterotrophic flagellate. J. Eukaryot. Microbiol. 40: 172-179.
Gray, M. W., Lang, B. F. & Burger, G. (2004) Mitochondria of protists. Annu. Rev. Genet. 38: 477-525.
Lang, B. F., Burger, G., OKelly, C. J., Cedergren, R., Golding, G. B., Lemieux, C., Sankoff, D., Turmel, M. & Gray, M. W. (1997) An ancestral mitochondrial DNA resembling a eubacterial genome in miniature. Nature 387: 493-497.
Lang, B. F., Gray, M.W., & Burger, G. (1999a) Mitochondrial genome evolution and the origin of eukaryotes. Annu. Rev. Genet. 33: 351-397.
Lang, B. F., Seif, E., Gray, M., OKelly, C. & Burger, G. (1999b) A comparative genomics approach to the evolution of the eukaryotes and their mitochondria. J. Eukaryot. Microbiol. 46: 320-326.
Lara, E., Chatzinotas, A., Simpson, A.G.B. (2006) Andalucia (gen. nov,): a new taxon for the deepest branch within jakobids (Jakobida; Excavata), based on morphological and molecular study of a new flagellate from soil. J. Eukaryot. Microbiol. 53: 112-120.
Nicholls, K.H. (1984) On the validity of Histiona aroides Pascher (Chrysophyceae?). Arch Protistenkd 128: 141-146
Mylnikov, A.P. (1984) The morphology and life cycle of Histiona aroides Pascher (Chrysophyta). Biologiia Vnutrennikh Vod: Informatsionnyi Biulleten 62: 16-19. [In Russian]
Mylnikov, A.P. (1989) The fine structure and systematic position of Histiona aroides (Bicoecales). Botaniheskei Zhurnal 74: 184--189 (in Russian).
OKelly, C. J. (1993) The jakobid flagellates: structural features of Jakoba, Reclinomonas and Histiona and implications for the early diversification of eukaryotes. J. Eukaryot. Microbiol. 40: 627--636.
OKelly, C.J. (1997) Ultrastructure of trophozoites, zoospores and cysts of Reclinomonas americana Flavin & Nerad, 1993 (Protista incertae sedis: Histionidae). Europ. J. Protistol. 33: 337-348.
OKelly, C.J. & Nerad, T.A. (1999) Malawimonas jakobiformis n. gen., n. sp. (Malawimonadidae n. fam.): A jakoba-like heterotrophic nanoflagellate with discoidal mitochondrial cristae. J. Eukaryot. Microbiol. 46: 522-531.
Pascher, A. (1943) Zur Klärung einiger gefärbter und farbloser Flagellaten und ihrer Einrichtungen zur Aufnahme animalischer Nahrung. Arch Protistenkd 96: 75-108.
Patterson, D.J. (1990) Jakoba libera (Ruinen, 1938), a heterotrophic flagellate from deep oceanic sediments. J. Mar. Biol. Ass. U.K. 70: 381-393.
Patterson, D.J., Vørs, N., Simpson, A.G.B. & O'Kelly, C.J. (2002) Residual and predatory heterotrophic flagellates. In: Lee, J.J., Leedale, G.F.& Bradbury P. (ed.) An illustrated guide to the protozoa, second edition. pp 1302-1328. Society of Protozoologists, Lawrence, Kansas.
Petersen, J.B. and Hansen, J.B. (1962) One some neuston organisms III. Svensk. Botanisk. Tidskrift 57: 293-305.
Rodríguez-Ezpeleta, N., Brinkmann, H., Burger, G., Roger, A.J., Gray, M.W., Philippe, H., Lang, B.F. (2007) Toward resolving the eukaryotic tree: The phylogenetic positions of jakobids and cercozoans. Curr. Biol. 17: 1420-1425.
Ruinen J. (1938) Notizen über Salzflagellaten. II. Über die Verbereitung der Salzflagellaten. Arch Protistenkd 90: 210-258.
Simpson, A.G.B. (2003) Cytoskeletal organisation phylogenetic affinities and systematics in the contentious taxon Excavata (Eukaryota). Int. J. Syst. Evol. Microbiol. 53: 1759-1777.
Simpson, A.G.B. & Patterson, D.J. (1999) The ultrastructure of Carpediemonas membranifera (Eukaryota) with reference to the excavate hypothesis. Europ. J. Protistol. 35: 353-370.
Simpson, A.G.B. & Patterson, D.J. (2001) On core jakobids and excavate taxa: the ultrastructure of Jakoba incarcerata. J. Eukaryot. Microbiol. 48: 480-492.
Simpson, A.G.B., Roger, A.J., Silberman, J.D., Leipe, D.D., Edgcomb, V.P., Jermiin, L.S., Patterson, D.J. & Sogin, M.L. (2002) Evolutionary history of early diverging eukaryotes: the excavate taxon Carpediemonas is a close relative of Giardia. Mol. Biol. Evol. 19: 1782-1791.
Simpson, A.G.B. and Roger, A.J. (2004) Excavata and the origin of amitochondriate eukaryotes. In Hirt, R. P. and Horner, D. S. (eds) Organelles, genomes and eukaryote phylogeny: an evolutionary synthesis in the age of genomics. pp. 27-53. CRC Press, London.
Simpson, A.G.B., Inagaki, Y., Roger, A.J., (2006) Comprehensive multigene phylogenies of excavate protists reveal the evolutionary positions of primitive eukaryotes. Mol. Biol. Evol. 23: 615-625.
Simpson, A.G.B., Perley, T. & Lara, E., (2008) Lateral transfer of the gene for a widely used marker, alpha tubulin, indicated by a multi-protein study of the phylogenetic position of Andalucia (Excavata). Mol. Phylogenet. Evol. 47: 366-377.
Yoon, H.S., and 9 others (2008) Broadly sampled multigene trees of eukaryotes. BMC Evol. Biol. 8: art. 14.
Yubuki, N., Inagaki, Y., Nakayama, T., Inouye, I., (2007) Ultrastructure and ribosomal RNA phylogeny of the free-living heterotrophic flagellate Dysnectes brevis n. gen., n. sp., a new member of the Fornicata. J. Eukaryot. Microbiol. 54: 191-200.
Title Illustrations

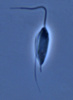
| Scientific Name | Jakoba libera |
|---|---|
| Comments | cell from ATCC strain 50422 |
| Source | Jakoba libera |
| Source Collection | Micro*scope |
| Image Use |
 This media file is licensed under the Creative Commons Attribution-ShareAlike License - Version 2.5. This media file is licensed under the Creative Commons Attribution-ShareAlike License - Version 2.5.
|
| Copyright | © |
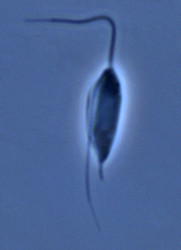
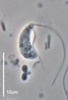
| Scientific Name | Andalucia incarcerata |
|---|---|
| Specimen Condition | Live Specimen |
| Image Use |
 This media file is licensed under the Creative Commons Attribution-NonCommercial License - Version 3.0. This media file is licensed under the Creative Commons Attribution-NonCommercial License - Version 3.0.
|
| Copyright |
© Alastair Simpson

|
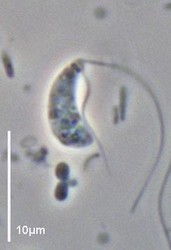
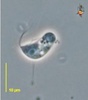
| Scientific Name | Reclinomonas americana |
|---|---|
| Source | Reclinomonas |
| Source Collection | Micro*scope |
| Image Use |
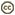 This media file is licensed under the Creative Commons Attribution-ShareAlike License - Version 2.5. This media file is licensed under the Creative Commons Attribution-ShareAlike License - Version 2.5.
|
| Copyright | © |
About This Page
This page is being developed as part of the Tree of Life Web Project Protist Diversity Workshop, co-sponsored by the Canadian Institute for Advanced Research (CIFAR) program in Integrated Microbial Biodiversity and the Tula Foundation.
Alastair Simpson

Dalhousie University, Halifax, Nova Scotia, Canada
Correspondence regarding this page should be directed to Alastair Simpson at
Page copyright © 2009 Alastair Simpson
 Page: Tree of Life
Jakobida.
Authored by
Alastair Simpson.
The TEXT of this page is licensed under the
Creative Commons Attribution-NonCommercial License - Version 3.0. Note that images and other media
featured on this page are each governed by their own license, and they may or may not be available
for reuse. Click on an image or a media link to access the media data window, which provides the
relevant licensing information. For the general terms and conditions of ToL material reuse and
redistribution, please see the Tree of Life Copyright
Policies.
Page: Tree of Life
Jakobida.
Authored by
Alastair Simpson.
The TEXT of this page is licensed under the
Creative Commons Attribution-NonCommercial License - Version 3.0. Note that images and other media
featured on this page are each governed by their own license, and they may or may not be available
for reuse. Click on an image or a media link to access the media data window, which provides the
relevant licensing information. For the general terms and conditions of ToL material reuse and
redistribution, please see the Tree of Life Copyright
Policies.
- First online 05 September 2008
- Content changed 05 September 2008
Citing this page:
Simpson, Alastair. 2008. Jakobida. Version 05 September 2008. http://tolweb.org/Jakobida/97407/2008.09.05 in The Tree of Life Web Project, http://tolweb.org/





 Go to quick links
Go to quick search
Go to navigation for this section of the ToL site
Go to detailed links for the ToL site
Go to quick links
Go to quick search
Go to navigation for this section of the ToL site
Go to detailed links for the ToL site