Eubacteria
True Bacteria


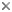
This tree diagram shows the relationships between several groups of organisms.
The root of the current tree connects the organisms featured in this tree to their containing group and the rest of the Tree of Life. The basal branching point in the tree represents the ancestor of the other groups in the tree. This ancestor diversified over time into several descendent subgroups, which are represented as internal nodes and terminal taxa to the right.

You can click on the root to travel down the Tree of Life all the way to the root of all Life, and you can click on the names of descendent subgroups to travel up the Tree of Life all the way to individual species.
For more information on ToL tree formatting, please see Interpreting the Tree or Classification. To learn more about phylogenetic trees, please visit our Phylogenetic Biology pages.
close boxReferences
Ahmad, S., A. Selvapandiyan, and R. K. Bhatnagar. 1999. A protein-based phylogenetic tree for Gram-positive bacteria derived from hrcA, a unique heat-shock regulatory gene. International Journal of Systematic Bacteriology 49:1387-1394.
Andersson, S. G. E., A. Zomorodipour, J. O. Andersson, T. Sicheritz-Ponten, U. C. M. Alsmark, R. M. Podowski, A. K. Naslund, A. S. Eriksson, H. H. Winkler, and C. G. Kurland. 1998. The genome sequence of Rickettsia prowazekii and the origin of mitochondria. Nature 396:133-140.
Balows, A., H.G. Träper, M. Dworkin, W. Harder, and K.-H. Schleifer (eds.). 1992. The Prokaryotes. A Handbook on the Biology of Bacteria: Ecophysiology, Isolation, Identification, Applications. Second edition, Volumes I-IV. Springer Verlag, New York.
Battistuzzi, F. U., A. Feijao, and A. B. Hedges. 2004. A genomic timescale of prokaryote evolution: insights into the origin of methanogenesis, phototrophy, and the colonization of land. BMC Evolutionary Biology 4:44-.
Battistuzzi, F. U. and A. B. Hedges. 2009. A major clade of prokaryotes with ancient adaptations to life on land. Molecular Biology and Evolution 26(2):335-343; doi:10.1093/molbev/msn247
Bern, M. and D. Goldberg. 2005. Automatic selection of representative proteins for bacterial phylogeny. BMC Evolutionary Biology 5:34-.
Boone, D. R., R.W. Castenholz, and G.M. Garrity. 2001. Bergey's Manual of Systematic Bacteriology. Springer, New York.
Brochier, C., E. Bapteste, D. Moreira, and H. Philippe. 2002. Eubacterial phylogeny based on translational apparatus proteins.
Trends in Genetics 18:1-5.
Brown, J. R. , C. J. Douady, M. J. Italia, W. E. Marshall, and M. J. Stanhope. 2001. Universal trees based on large combined protein sequence data sets. Nature Genetics 28:281-285.
Bustard, K. and R. S. Gupta. 1997. The sequences of heat shock protein 40 (DnaJ) homologs provide evidence for a close evolutionary relationship between the Deinococcus-Thermus group and cyanobacteria. Journal of Molecular Evolution 45:193-205.
Castro, H. F., N. H. Williams, and A. Ogram. 2000. Phylogeny of sulfate-reducing bacteria. FEMS Microbiology Ecology 31:1-9.
Cavalier-Smith, T. 2002. The neomuran origin of archaebacteria, the negibacterial root of the universal tree and bacterial megaclassification. International Journal of Systematic and Evolutionay Microbiology 52:7-76.
Coenye, T. and P. Vandamme. 2004. A genomic perspective on the relationship between the Aquificales and the epsilon-Proteobacteria. Syst Appl. Microbiol. 27:313-322.
Daubin, V., M. Gouy, and G. Perriere. 2002. A phylogenomic approach to bacterial phylogeny: Evidence of a core of genes sharing a common history. Genome Research 2(7):1080-1090.
DeLong E. F. and N. R. Pace. 2001. Environmental diversity of Bacteria and Archaea. Systematic Biology 50:470-478.
Eisen, J. A. 1995. The RecA protein as a model molecule for molecular systematic studies of bacteria: Comparison of trees of RecAs and 16S rRNAs from the same species. Journal of Molecular Evolution 41:1105-1123. Molecular Biology and Evolution 21(9):1643-1660.
Emelyanov, V. V. and B. V. Sinitsyn. 1999. A groE-based phylogenetic analysis shows very close evolutionary relationship between mitochondria and Rickettsia. Russian Journal of Genetics 35:618-627.
Esser, C., N. Ahmadinejad, C. Wiegand, C. Rotte, F. Sebastiani, G. Gelius-Dietrich, K. Henze, E. Kretschmann, E. Richly, D. Leister, D. Bryant, M. A. Steel, P. J. Lockhart, D. Penny and W. Martin. 2004. A genome phylogeny for mitochondria among alpha-proteobacteria and a predominantly eubacterial ancestry of yeast nuclear genes.
Fox, G. E., E. Stackebrandt, R. B. Hespell, J. Gibson, J. Maniloff, T. A. Dyer, R. S. Wolfe, W. E. Balch, R. S. Tanner, L. J. Magrum, L. B. Zablen, R. Blakemore, R. Gupta, L. Bonen, B. J. Lewis, D. A. Stahl, K. R. Luehrsen, K. N. Chen, and C. R. Woese. 1980. The phylogeny of prokaryotes. Science 209:457-463.
Garrity, G. M., J. A. Bell, and T. G. Lilburn. 2004. Taxonomic Outline of the Prokaryotes. Bergey's Manual of Systematic Bacteriology, Second Edition. Release 5.0.
Gray, M. W., G. Burger, and B. F. Lang. 1999. Mitochondrial evolution. Science 283:1476-1481.
Griffiths, E. and R. S. Gupta. 2004. Signature sequences in diverse proteins provide evidence for the late divergence of the Order Aquificales. Int Microbiol. 7:41-52.
Gruber, T. M. and D. A. Bryant. 1997. Molecular systematic studies of eubacteria, using sigma(70)-type sigma factors of group 1 and group 2. Journal of Bacteriology 179:1734-1747.
Gupta, R. S. 1997. Protein phylogenies and signature sequences: Evolutionary relationships within prokaryotes and between prokaryotes and eukaryotes. Antonie van Leeuwenhoek International Journal of General and Molecular Microbiology 72:49-61.
Gupta, R. S. 1998. Protein phylogenies and signature sequences: A reappraisal of evolutionary relationships among archaebacteria, eubacteria, and eukaryotes. Microbiology and Molecular Biology Reviews 62:1435-1491.
Gupta, R. S. 2000. The phylogeny of proteobacteria: relationships to other eubacterial phyla and eukaryotes. FEMS Microbiology Reviews 24(4):367-402.
Gupta, R. S. 2004. The phylogeny and signature sequences characteristics of Fibrobacteres, Chlorobi, and Bacteroidetes. Critical Reviews in Microbiology 30(2):123-143.
Gupta, R.S., K. Bustard, M. Falah, D. Singh. 1997. Sequencing of heat shock protein 70 (DnaK) homologs from Deinococcus proteolyticus and Thermomicrobium roseum and their integration in a protein-based phylogeny of prokaryotes. Journal of Bacteriology 179:345-357.
Gupta, R.S. and E. Griffiths. 2002. Critical issues in bacterial phylogeny. Theoretical Population Biology 61(4):423-434.
Gupta, R. S., T. Mukhtar, and B. Singh. 1999. Evolutionary relationships among photosynthetic prokaryotes (Heliobacterium chlorum, Chloroflexus aurantiacus, cyanobacteria, Chlorobium tepidum and proteobacteria): implications regarding the origin of photosynthesis. Molecular Microbiology 32:893-906.
Huang, W. M. 1996. Bacterial diversity based on type II DNA topoisomerase genes. Annual Review of Genetics 30:79-107.
Huang, Y. P. and J. Ito. 1999. DNA polymerase C of the thermophilic bacterium Thermus aquaticus: Classification and phylogenetic analysis of the family C DNA polymerases. Journal of Molecular Evolution 48:756-769.
Hugenholtz, P., B. M. Goebel, and N. R. Pace. 1998. Impact of culture-independent studies on the emerging phylogenetic view of bacterial diversity. Journal of Bacteriology 180:4765-4774.
Kunisawa, T. 2006. Dichotomy of major bacterial phyla inferred from gene arrangement comparisons. J. of Theor. Biol. 239:367-375.
Lawrence, J. G. and H. Ochman. 1998. Molecular archaeology of the Escherichia coli genome. Proceedings of the National Academy of Sciences of the United States of America 95:9413-9417.
Liu, R. and H. Ochman. 2007. Stepwise formation of the bacterial flagellar system. Proceedings of the National Academy of Sciences of the United States of America 104(17):7116-7121.
Ludwig, W., J. Neumaier, N. Klugbauer, E. Brockmann, C. Roller, S. Jilg, K. Reetz, I. Schachtner, A. Ludvigsen, M. Bachleitner, U. Fischer, and K. H. Schleifer. 1993. Phylogenetic relationships of Bacteria based on comparative sequence analysis of elongation factor Tu and ATP-synthase beta-subunit genes. Antonie van Leeuwenhoek International Journal of General and Molecular Microbiology 64:285-305.
Ludwig, W., O. Strunk, S. Klugbauer, N. Klugbauer, M. Weizenegger , J. Neumaier, M. Bachleitner, and K. H. Schleifer. 1998. Bacterial phylogeny based on comparative sequence analysis. Electrophoresis 19:554-568.
Moran, N. and P. Baumann. 1994. Phylogenetics of cytoplasmically inherited microorganisms of arthropods. Trends in Ecology and Evolution 9:15-20.
Ochman, H., S. Elwyn, and N. A. Moran. 1999. Calibrating bacterial evolution. Proceedings of the National Academy of Sciences of the United States of America 96:12638-12643.
Olsen, G. J. and C. R. Woese. 1993. Ribosomal RNA: a key to phylogeny. FASEB Journal 7:113-23.
Olsen, G. J., C. R. Woese, and R. Overbeek. 1994. The winds of (evolutionary) change: breathing new life into microbiology. Journal of Bacteriology 176:1-6.
Pace, N. R. 1997. A molecular view of microbial diversity and the biosphere. Science 276:734-740.
Pace, N. R. 1999. Microbial ecology and diversity. ASM News 65:328-333.
Pierson, B. K. 1994. The emergence, diversification, and role of photosynthetic bacteria. Pages 161-180 in Early Life on Earth, Nobel Symposium No. 84 (Bengtson, S., ed.). Columbia University Press, New York.
Rappé, M. S. and S. J. Giovannoni. 2003. The uncultured microbial majority. Annual Review of Microbiology 57:369-394.
Sapp, J. 2005) Microbial Phylogeny and Evolution: Concepts and Controversies. Oxford University Press, New York.
Sicheritz-Ponten, T., C. G. Kurland, and S. G. E. Andersson. 1998. A phylogenetic analysis of the cytochrome b and cytochrome c oxidase I genes supports an origin of mitochondria from within the Rickettsiaceae. Biochimica et Biophysica Acta Bioenergetics 1365:545-551.
Staley, J. T. and J. J. Gosink. 1999. Poles apart: Biodiversity and biogeography of sea ice bacteria. Annual Review of Microbiology 53:189-215.
Stetter, K. O. 1996. Hypterthermophilic procaryotes. FEMS Microbiology Reviews 18:149-158.
Teichmann, S. A. and G. Mitchison. 1999. Is there phylogenetic signal in prokaryote proteins? Journal of Molecular Evolution 49:98-107.
Viale, A. M. and A. K. Arakaki. 1994. The chaperone connection to the origins of the eukaryotic organelles. FEBS Letters 341:146-151.
Viale, A. M., A. K. Arakaki, F. C. Soncini, and R. G. Ferreyra. 1994. Evolutionary relationships among eubacterial groups as inferred from GroEL (chaperonin) sequence comparison. International Journal of Systematic Bacteriology 44:527-533.
Wagner, M. and M. Horn. 2006. The Planctomycetes, Verrucomicrobia, Chlamydiae and sister phyla comprise a superphylum with biotechnological and medical relevance. Curr. Opin. Biotech. 17:241-249.
Woese, C. R. 1987. Bacterial evolution. Microbiological Reviews 51:221-271.
Wolf, Y. I., I. B. Rogozin, N. V. Grishin, R. L. Tatusov, and E. V. Koonin. 2001. Genome trees constructed using five different approaches suggest new major bacterial clades. BMC Evolutionary Biology 1:8-.
Yang, S., R. F. Doolittle, and P. E. Bourne. 2005. Phylogeny determined by protein domain content. Proceedings of the National Academy of Science USA 102(2):373-378 .
Information on the Internet
- Phylogenomics: A Genome Level Approach to Assembling the Bacterial Branches of the Tree of Life. The Institute for Genomic Research.
- The Prokaryotes. An evolving electronic resource for the microbiological community.
- Introduction to the Bacteria. UCMP Berkeley.
- MicrobeWorld. American Society of Microbiology
- Microbes.info. The Microbiology information portal.
- Virtual Museum of Bacteria
- Digital Learning Center for Microbial Ecology. A science education project at Michigan State University.
- Bacteriology 303. Study aids and background readings. University of Wisconsin - Madison.
- The Microbial World: Profiles of Microorganism. An educational resource covering bacteria, fungi, viruses, biological control, environmental microbiology, disease of plants, humans and animals. Produced by Jim Deacon, Institute of Cell and Molecular Biology, and Biology Teaching Organisation, University of Edinburgh.
- Stalking the Mysterious Microbe. A kid-oriented site from the American Society for Microbiology.
- Bugs in the News!. Interesting facts about microbes compiled by John C. (Jack) Brown.
- The Microbiology Network. A communication resource for the microbiologist.
- List of Bacterial names with Standing in Nomenclature. J. P. Euzéby.
- TIGR Microbial Database. A listing of microbial genomes and chromosomes completed and in progress.
- Bergey's Manual of Systematic Bacteriology, second edition. The taxonomic outline for the 5 volumes, including all species, type strains, and 16S rRNA sequence data can be downloaded from this site.
- Bacterial Nomenclature Up-To-Date. DSMZ-Deutsche Sammlung von Mikro-organismen und Zellkulturen GmbH, Braunschweig, Germany.
- The"Bad Bug Book". Foodborne Pathogenic Microorganisms and Natural Toxins Handbook. U.S. Food & Drug Administration Center for Food Safety & Applied Nutrition.
- Cells Alive!. Videos and images of bacteria and other organisms.
- Public Health Image Library. Centers for Disease Control and Prevention (CDC).
Title Illustrations

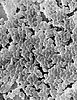
| Scientific Name | Enterococcus |
|---|---|
| Comments | Scanning Electron Micrograph of Enterococcus |
| Creator | CDC/Janice Carr |
| Specimen Condition | Dead Specimen |
| Source Collection | Public Health Image Library (Centers for Disease Control) |
| Scientific Name | Pseudomonas aeruginosa |
|---|---|
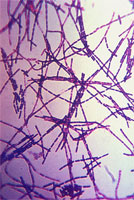
| Comments | Scanning Electron Micrograph of Pseudomonas aeruginosa |
| Creator | CDC/Janice Carr |
| Specimen Condition | Dead Specimen |
| Source Collection | Public Health Image Library (Centers for Disease Control) |
| Scientific Name | Bacillus anthracis |
|---|---|
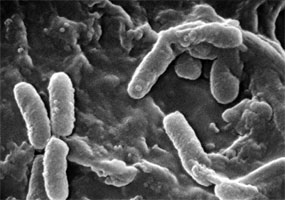
| Comments | A photomicrograph of the Anthrax bacterium, Bacillus anthracis, using Gram stain technique |
| Specimen Condition | Dead Specimen |
| Source Collection | Public Health Image Library (Centers for Disease Control) |
About This Page
Page copyright © 2005
 Page: Tree of Life
Eubacteria. True Bacteria.
The TEXT of this page is licensed under the
Creative Commons Attribution-NonCommercial License - Version 3.0. Note that images and other media
featured on this page are each governed by their own license, and they may or may not be available
for reuse. Click on an image or a media link to access the media data window, which provides the
relevant licensing information. For the general terms and conditions of ToL material reuse and
redistribution, please see the Tree of Life Copyright
Policies.
Page: Tree of Life
Eubacteria. True Bacteria.
The TEXT of this page is licensed under the
Creative Commons Attribution-NonCommercial License - Version 3.0. Note that images and other media
featured on this page are each governed by their own license, and they may or may not be available
for reuse. Click on an image or a media link to access the media data window, which provides the
relevant licensing information. For the general terms and conditions of ToL material reuse and
redistribution, please see the Tree of Life Copyright
Policies.
- Content changed 10 March 2006
Citing this page:
Tree of Life Web Project. 2006. Eubacteria. True Bacteria. Version 10 March 2006 (temporary). http://tolweb.org/Eubacteria/2/2006.03.10 in The Tree of Life Web Project, http://tolweb.org/







 Go to quick links
Go to quick search
Go to navigation for this section of the ToL site
Go to detailed links for the ToL site
Go to quick links
Go to quick search
Go to navigation for this section of the ToL site
Go to detailed links for the ToL site